Volume 10, Issue 5 (September & October 2019)
BCN 2019, 10(5): 485-498 |
Back to browse issues page
Download citation:
BibTeX | RIS | EndNote | Medlars | ProCite | Reference Manager | RefWorks
Send citation to:



BibTeX | RIS | EndNote | Medlars | ProCite | Reference Manager | RefWorks
Send citation to:
Grüßer L, Blaumeiser-Debarry R, Rossaint R, Krings M, Kremer B, Höllig A et al . A 6-step Approach to Gain Higher Quality Results From Organotypic Hippocampal Brain Slices in a Traumatic Brain Injury Model. BCN 2019; 10 (5) :485-498
URL: http://bcn.iums.ac.ir/article-1-1019-en.html
URL: http://bcn.iums.ac.ir/article-1-1019-en.html
Linda Grüßer *1 

 , Rosmarie Blaumeiser-Debarry1
, Rosmarie Blaumeiser-Debarry1 
 , Rolf Rossaint1
, Rolf Rossaint1 

 , Matthias Krings1
, Matthias Krings1 
 , Benedikt Kremer2
, Benedikt Kremer2 
 , Anke Höllig3
, Anke Höllig3 

 , Mark Coburn1
, Mark Coburn1 




 , Rosmarie Blaumeiser-Debarry1
, Rosmarie Blaumeiser-Debarry1 
 , Rolf Rossaint1
, Rolf Rossaint1 

 , Matthias Krings1
, Matthias Krings1 
 , Benedikt Kremer2
, Benedikt Kremer2 
 , Anke Höllig3
, Anke Höllig3 

 , Mark Coburn1
, Mark Coburn1 


1- Department of Anesthesiology, RWTH Aachen University Hospital, Aachen, Germany.
2- Department of Neurosurgery, RWTH Aachen University Hospital, Pauwelsstr. 30, 52074 Aachen, Germany
3- Department of Neurosurgery, RWTH Aachen University Hospital, Aachen, Germany.
2- Department of Neurosurgery, RWTH Aachen University Hospital, Pauwelsstr. 30, 52074 Aachen, Germany
3- Department of Neurosurgery, RWTH Aachen University Hospital, Aachen, Germany.
Keywords: Organotypic hippocampal brain slices, In vitro model, Traumatic Brain Injury (TBI), Propidium Iodide (PI), Eligibility, Framework
Full-Text [PDF 1914 kb]
| Abstract (HTML)
2.1.5.1. Confirmation
In the first step, it is recommended to verify that each picture taken shows only one slice, amounts to the pixels of this slice, and thus fully shows its cell damage. We suggest enhancing the contrast of each image captured (only for better visibility, pixel counts are determined as explained above) (Figure 2.1). This procedure allows for the identification of a second slice in the picture and eventually the blackening of this area. Further, a thorough evaluation of each photograph is advantageous to detect small PI-clots or fusels. Finally, we are sure that each image taken before the experiment is matched with the correct one at later time-points of assessment (Figure 2.1, Example image step 1).
2.1.5.2. Identification of pre-damaged slices at baseline assessment and exclusion
Other researchers have pointed to the fact, that they excluded slices from their experiments if they were damaged at baseline assessment or showed high levels of PI fluorescence (Adamchik et al., 2000; Harris et al., 2013; Krings et al., 2016; Miller et al., 2015). However, no details were provided, and in many papers, we did not find any information that slices were excluded. As inserts are moved for medium changes and are exposed to temperature fluctuations, pre-damage to the slice at baseline assessment is a common event. Moreover, preparation techniques and timing may still improve within the course of the experiment.
It is crucial to avoid pre-damaged slices entering into the experiment, as their trauma intensity may falsify the results after the incubation time. Since it is unknown whether a higher amount of injured cells at baseline assessment affect or even exponentially increase the development of secondary injury, only minimally pre-damaged slices should be considered in the experiment. An objective threshold should be defined to avoid bias in identification and outsourcing of these slices. In our research, if baseline image histograms presented more than 1000 pixels, they would be sorted out as they were considered to be pre-damaged (Figure 2.2, Example image of step 2).
2.1.5.3. Preparation margin error
Motor skills develop over training sessions and probably through different stages (Luft & Buitrago, 2005). Similarly, preparation techniques improve when regularly dissecting the hippocampi. A typical feature we observed, especially within the first rounds of prepared slices, was a prominent margin due to-as suggested here-errors in preparation. Even though preparation steps were always strictly followed, some of the slices presented with noticeable frames of PI-detectable cells. The pictures taken at baseline assessment did not necessarily exceed an integral of >1000 pixels.
After an incubation period of two hours, however, PI fluorescence was highly visible, and thus slices with such a prominent margin distorted the outcome of the affected groups. In order to objectively decide on an allowable error in preparation and an average damage which would inevitably occur, the following threshold was applied: if the PI-detectable cell death in a margin exceeded >250 pixels, this margin was considered to be prominent enough to be an error stemming from errors in the preparation and its pixels were not added to the full integration of pixels in this picture (Figure 2.3). We took the same approach for slices presenting with a “one-celled” margin, which was observed in only two experimental series (Figure 2.3, Example image of step 3).
2.1.5.4. Morphological peculiarities
In this study, we noticed several morphological peculiarities in varying degrees. These peculiarities include what we describe as “holes”, “frayed margins”, and “emigrated cells” (Figure 2.4). A closer investigation of these phenomena is suggested for future studies, as we do not know why they occurred and cannot evaluate whether electrophysiological consequences are entailed. We observed that “holes” often went in line with “frayed margins” and “emigrated cells”.
However, these features also occurred independently. Additionally, by enhancing the contrast of the image, we noticed that emigrated cells even existed at baseline assessment, but this was not always detectable by PI. Excluding all slices with a specific morphology would significantly decrease the number of slices available in particular groups. Therefore, we decided to refer to step 1: if slices with peculiarities were presented with <1000 pixels at baseline assessment, they would be taken into consideration for the experiment (Figure 2.4, Example image of step 4).
2.1.5.5. Variation of trauma intensities
During the experiment, we observed that trauma intensities varied over time and that some experimental series, even within one group presented with different trauma severities despite the same dropping height was ensured (Figure 2.5). Thus, we saw the necessity to define a scope of trauma strength. A boxplot analysis was performed for the trauma control group, and extreme outliers were identified (n=2) and excluded. The new mean value of the trauma control group plus two times the standard deviation served as an upper threshold for all the other trauma groups.
Concerning trauma intensities and depending on where the pin hits the slice, more or less tissue can be injured by secondary injury; that is, if the pin hits the slice close to the margin there is less hippocampus to be affected in the following time. Also the shapes of the hippocampi- as we experienced- vary where we found it is unlikely to work with remarkably similar contoured slices. Hence, our approach was to consider all slices except for those that were not hit by the pin (n=7).
In our experiment, we identified several borderline cases, mostly in the intervention group presenting with the least secondary injury, where it appeared to be difficult to make a clear decision. Future researchers are recommended to set a defined size before the experiment that has to be considered for inclusion criteria, including the hippocampus itself (which is also of importance for the non-trauma groups), the pin’s impression, and the area of secondary injury (Figure 2.5, Example image for step 5).
2.1.5.6. Susceptibility of non-trauma groups
We noticed that it is crucial to adjust the focus of the camera in taking pictures, particularly in the non-trauma groups, where trauma intensities are minimal. Blurry images can present with a lower amount of greyscale pixels as the majority of pixels are evaluated as background fluorescence. Hence, in a few cases, pre-damaged slices may not be excluded by the 1000-pixels threshold and then present with excessive greyscale values after the incubation time. Also- the other way round- a misleadingly low number of pixels is possible when the picture was taken after the incubation time is blurry.
To prevent biases by manually evaluating the sharpness of hundreds of pictures, we first excluded all images in the non-trauma groups that had lower pixel numbers after the incubation period than at baseline assessment. We subjectively excluded 11 slices due to their size as, for example, they were too small to be considered a hippocampal slice. In the future, a defined size of the hippocampus should be set before the experiment. In the next step, we performed a boxplot-analysis in each non-trauma group to exclude outliers. Thus, we objectively excluded two kinds of slices: the slices presenting with excessive trauma (as the pictures showing the pre-damage at baseline assessment were blurry), and the slices damaged due to careless handling or temperature change (Figure 2-6, Example image of step 6).
3. Results
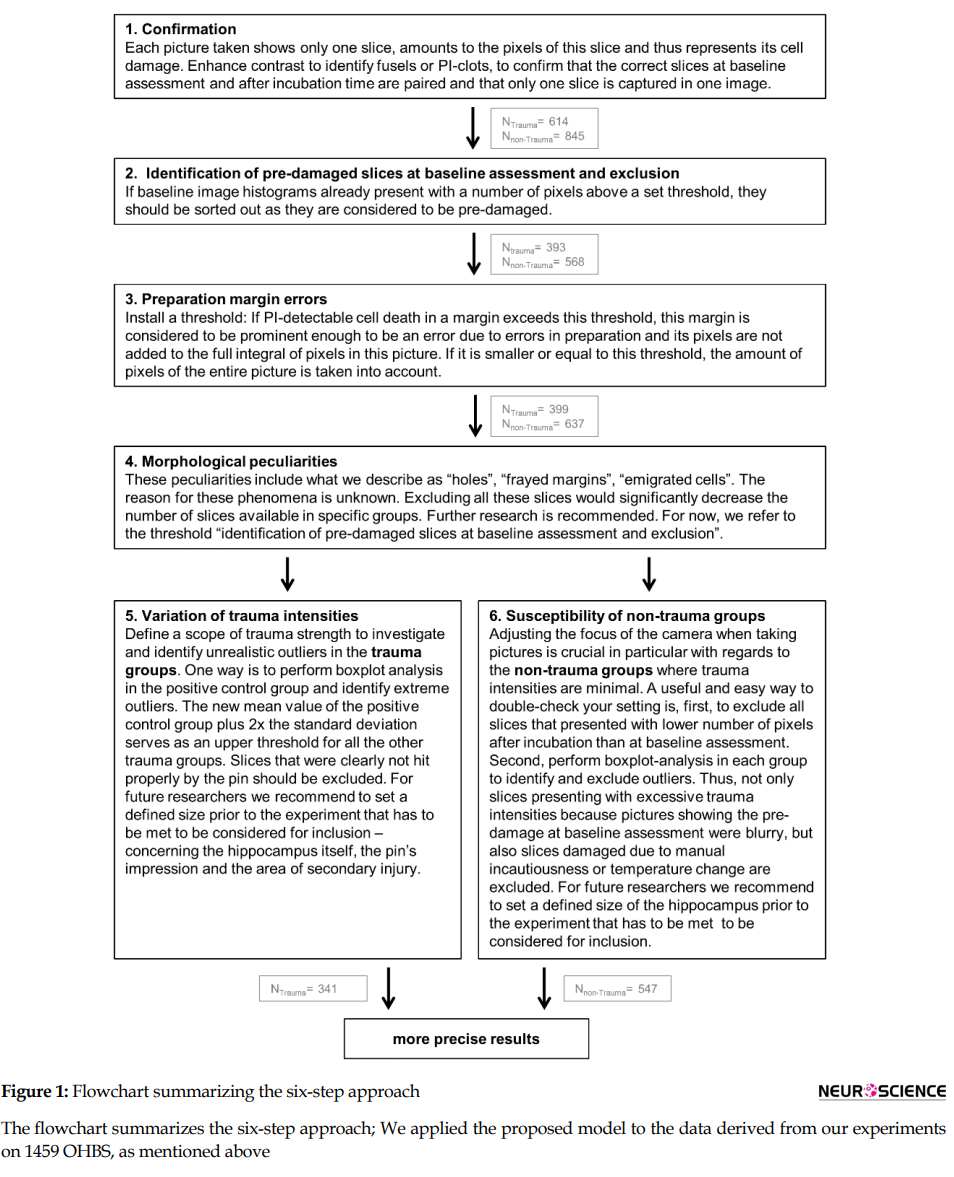
We proposed a six-step approach to enhance comparability and applied the proposed model to the data derived from our experiments on 1459 OHBS, as mentioned above (Figure 1). Then we presented exemplary results that demonstrated how the outcome could differ when a measure was applied vs. when it was not.
After the first checkpoint (step 1) (confirmation), 1459 slices were assessed for pre-damage (step 2). As Figure 3.1 shows, the difference between two non-trauma groups seems to be significant at first glance; yet, when it is ascertained that baseline slices in both groups were presented with similar PI-detectable trauma intensities, this difference did not turn out to be significant anymore (P=0.004 vs. P=0.716; ∆ intervention 2: 0.27; ∆ intervention 3: 0.56). In the next step, the slices were screened for preparatory margins (step 3). Figure 3.2 demonstrates the difference in results when preparation errors are considered vs. when they are not. Misleadingly, the non-trauma/intervention group 2 would have been considered significantly more injured than the non-trauma/intervention group 1 (P=0.004 vs. P=0.444; ∆ intervention 1: 0.02 ∆ intervention 2: 0.24).
Without applying this measure, a difference between groups would have mistakenly been attributed to the specific interventions. By subtracting margins stemming from errors in the preparatory process, more slices met the baseline assessment threshold and were considered for the next steps (393 slices after step 2 vs. 399 slices after step 3 in the trauma groups and 568 vs. 637 slices in the non-trauma groups). Morphological peculiarities, as described in step 4 require further investigation. Step 5 aims to set a scope concerning trauma intensities in the trauma groups.
According to Figure 3.3, without a set scope, the trauma/intervention group 3 would misleadingly have been considered to generate a neuro-harming effect in comparison to the trauma/intervention group 2 (P<0.001 vs. P=0.304; ∆ intervention 2: 0.02; ∆ intervention 3: 0.31). Identifying and excluding outliers in the non-trauma groups (step 6) revealed that the difference between the non-trauma/no intervention group and the non-trauma/intervention group 3 was not significant (P=0.016 vs. P=0.075; ∆ no intervention: 0.24; ∆ intervention 3: 0.40) (Figure 3.4).
After the six-step approach, the number of slices was narrowed down to 547 pictures in the non-trauma groups and 341 photos in the trauma groups (Figure 1). Therefore, 61% of the slices met the proposed inclusion criteria (Figure 3; results demonstrating more precise findings).
4. Discussion
Working with OHBS in a TBI-experiment, we noticed that several factors might influence the results. To our knowledge, detailed objective manners to generate more comparable results in such a susceptible set-up have not been discussed so far. The proposed model aims to set precise requirements to lower the possibility of pre-damaged slices entering into the experiment and outliers twisting the final results. This methodological approach provides the first framework to cope with the broad variation observed in these models. Our exemplary results show that more precise findings are ensured by setting equal requirements for all slices.
In vitro models provide a useful tool to scrutinize processes and treatment options in the initial step. Between dissociated cell cultures and the in vivo state, OHBS presents a potent alternative (Humpel, 2015; Li et al., 2016; Morrison, Saatman, Meaney, & McIntosh, 1998; Noraberg et al., 2005). The hippocampal circuits in organotypic brain slices, which are usually derived from early postnatal animals, are found to be similar to the in vivo state; however, some differences have been detected (Gähwiler et al., 1997; Gogolla, Galimberti, DePaola, & Caroni, 2006).
Of course, variables like perfusion, intracranial pressure, or the blood-brain barrier have not yet reflected in organotypic brain slice models, and the use of adult donor animals for organotypic brain slices has been scarce (Humpel, 2015; Li et al., 2016). Nonetheless, slices have remained 100-150 µm thick with a three-dimensional organization of connectivity (Gähwiler et al., 1997; Gogolla et al., 2006; Li et al., 2016; Stoppini et al., 1991). Notably, outcomes of studies working with OHBS have successfully been reproduced in in vivo models demonstrating the potential of this setup. For example, argon and xenon displayed a neuroprotective profile in OHBS models which could be substantiated in in vivo studies (Brücken et al., 2013; Coburn et al., 2008; Harris et al., 2013; Loetscher et al., 2009; Zhuang et al., 2012).
An important implication of research with organotypic brain slices is the reduction of the number of animal experiments (Humpel, 2015; Li et al., 2016). Further, in in vitro models of CNS injury, the associated costs are lower (Morrison et al., 1998). In their reviews on brain slice models, Noraberg et al. concluded that organotypic brain slices emerged to be an increasingly popular tool to investigate neurological diseases. Cho et al. asserted that through advancements of brain slices and technological innovations in this field now a tremendous potential exists to address questions quicker (Cho et al., 2007; Noraberg et al., 2005).
To our knowledge, precise and objective procedures to make results more comparable within one experiment and among different studies have not been discussed in detail yet. Retrospectively, Guy et al. method of determining slice thickness as a parameter for OHBS health might present a useful additional measure (Guy, Rupert, Sandberg, & Weber, 2011). In their review, Humpel et al. generally recommended withdrawing thick and not flattened slices. An outgrowth of cells from the edge of the living slices and a change in color to transparent grey color in the first week are criteria described for ‘good’ slices (Humpel, 2015).
De Simoni et al. pointed the difficulty to identify pyramidal cell layers in an unhealthy slice (De Simoni & Yu, 2006). Also, Gogolla et al. recommended not to culture slices with a lesioned CA1 region or a lesioned/detached dentate gyrus but only slices ‘with nice cell layers in the dentate gyrus and CA1-3 and smooth margins’ seems to be a reasonable approach to be considered for future researchers (Gogolla et al., 2006). Our model mainly relied on PI-detectable cell death, but other cell-specific markers than PI can be used (Humpel, 2015).
Experimental papers often do not report about slice selection. Some researchers have pointed to difficulties as variation in PI-uptake, incomplete layers or pre-damage, but did not explain objective criteria concerning how or why brain slices were excluded or included (Adamchik et al., 2000; Harris et al., 2013; Krings et al., 2016; Li et al., 2016; Miller et al., 2015). For clinical trials, participants must meet specific standards, fulfill inclusion criteria, or be prohibited from participating due to exclusion criteria (National Institutes of Health, 2016). This method neither protects the participants’ safety nor ensures that the subsequent results are more comparable as the study population is more alike.
Within our TBI-experiment in OHBS, we found that results differ when hippocampal brain slices undergo specific assessments for eligibility and that hippocampal brain slices can present with pre-damage or specific peculiarities and identified factors that might affect the outcome.
In general, researchers should work as cautiously as possible to avoid imprecise preparation, temperature changes for a long time, accidental shaking, or changes in the experimental process. Although all precautions may be observed, organizational challenges concerning timing, the microscope setting, or the medium may occur. Fluctuations in the severity of trauma may distort the outcome. Although the reasons are not clear, our results show that hippocampal brain slices do not necessarily present an unconditionally homogenous group. Thus, eligibility criteria for OHBS present a plausible conclusion.
To avoid bias by manually evaluating slices, the criteria set should be as objective as possible. If slices are uniform and interventions are consistent, the eligibility criteria do not interfere with the outcome. If any inconsistencies occur, the applied measures increase the chances of more precise conclusions to be drawn. In our experiment, for example, a considerable amount of data were estimated unusable as the slices were considered pre-damaged. Since this threshold was chosen arbitrarily, we could thus ensure that equal requirements were set for all experimental series.
Our results demonstrate that outcomes in a TBI experiment with OHBS should be critically analyzed as inhomogeneity may occur. Altogether, the proposed six steps are designed to consider possible errors; they also help to contextualize the findings and make results more comparable. In the future, enhanced comparability may decrease the number of animals needed.
This framework presents a first attempt to enhance comparability in an in vitro model of TBI in OHBS. Further research is recommended to confirm and further develop this model in similar studies. It might improve the quality of the results if the general recommendations listed above were specified and also considered in the proposed approach because of PI-detectable cell death, and the proposed measures might not be sufficient.
Some challenges that could not be adequately addressed would remain. First, the proposed thresholds are chosen arbitrarily, and future researchers are recommended to find an analytical solution of what should be considered ineligible, i.e. pre-damage.
Another disadvantage might be the visible preparatory margins with less than 250 pixels could still enter the experiment. As the margins had to be identified and blackened manually, this represents a subjective component and remains a potential source of error. We aimed to blacken only areas of cell death, possibly stemming from errors in the preparation process as the morphological peculiarities and their origin are unknown. However, it is not always clear whether a specific area should be considered as a “preparatory error”, a “frayed margin”, a “hole”, or as “emigrated cells”. For further studies, we would also recommend to regularly allocate prepared slices in one series to all groups to distribute potential errors equally.
Despite all measures taken, the slices that were too small to be considered as a hippocampus or some slices which were not hit by the pin had to be identified. We tried to minimize the number of slices excluded manually, so this problem presented a subjective choice, and we recommend defining inclusion criteria concerning the size of the hippocampi and the trauma area for future research. This suggestion seems particularly useful, as the trauma produced by the pin in our experiment, does not always resemble a proper circle, as are observable in images shown in other papers (Adamchik et al., 2000; Roehl et al., 2010). It is unclear whether this is typical as the injury spreads non-circularly throughout the slice depending on differences in susceptibility of cells or is caused by other factors, as an oblique fall.
Although the framework was constructed to avoid future errors, there may be cases that were not fixed. For example, our measures to exclude the most blurry pictures might not be sufficient to identify them. We included all slices and applied the measures proposed; however, numerical thresholds, i.e. morphological completeness, cannot be identified. Nevertheless, we sorted out a considerable amount of data and when the inclusion criteria failed to meet. This model presented a first approach to make the research with hippocampal brain slices in an in vitro model of TBI more comparable.
5. Conclusion
Several factors may influence experimental outcomes in a TBI-experiment in OHBS. To achieve the most precise result, we should set checkpoints, define pre-damaged slices, and restrain from entering into the experiment. Besides, we should detect unrealistic outliers and eventually exclude them. We presented objective approaches in our framework and included thresholds and boxplot-analyses. Our exemplary results demonstrate more comparable findings. We recommend future researchers to confirm and develop further this model and to scrutinize morphological peculiarities, their origin, and their potential impact on cell survival and electrophysiological functioning.
Ethical Considerations
Compliance with ethical guidelines
All ethical principles were considered in this article.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Authors' contributions
All authors contributed in preparing this article.
Conflict of interest
The authors declared no conflict of interest.
Acknowledgments
We thank Prof. Joachim Weis, head of the Department of Neuropathology, RWTH Aachen University Hospital, Aachen, Germany, and his team for their support in the culturing of the slices.
References
Full-Text:
1. Introduction
Brain cells have been studied in cultures for over 100 years (Ross Granville Harrison, 1910; Rose G. Harrison, Greenman, Mall, & Jackson, 1907; Hogue, 1952; Humpel, 2015; Millet & Gillette, 2012). Over time, different methods were developed to preserve thin brain slices and keep them viable for a longer time (Cho, Wood, & Bowlby, 2007; Gähwiler, 1981; Humpel, 2015; Stoppini, Buchs, & Muller, 1991). Slice cultures can be prepared from a variety of brain regions; however, hippocampal slice cultures are used most frequently and replicate many aspects of the in vivo state, making possible investigations into mechanisms of brain synapses (Bahr et al., 1995; Cho et al., 2007; Finley et al., 2004; Gähwiler, Capogna, Debanne, McKinney, & Thompson, 1997; Li, Han, & Wang, 2016; Noraberg et al., 2005).
Given that the hippocampal slice set-up also provides decent experimental access and allows for detailed regulation of the extracellular environment, it is not surprising that these systems have been widely used to study neurogenesis or to investigate diseases such as Alzheimer and stroke (Cho et al., 2007; Noraberg et al., 2005).
Several researchers utilized organotypic hippocampal slice cultures to scrutinize processes and possible treatment approaches for Traumatic Brain Injury (TBI) and described different methods in this regard (Adamchik, Frantseva, Weisspapir, Carlen, & Perez Velazquez, 2000; Coburn, Maze, & Franks, 2008; Di Pietro et al., 2010; Di Pietro et al., 2013; Harris et al., 2013; Hughes, Silva, Ahmed, Shreiber, & Morrison, 2014; Krings, Höllig, Grüsser, Rossaint, & Coburn, 2016; Loetscher et al., 2009; Miller et al., 2015; Morrison, Cater, Benham, & Sundstrom, 2006; Morrison et al., 2003; Roehl et al., 2010; Rossaint et al., 2009; Schoeler et al., 2012; Smith et al., 2016; Vogel et al., 2016).
Morrison et al. described an injury device that stretched the hippocampal brain slices equi-biaxially to simulate brain deformation through TBI (Morrison et al., 2006). Di Pietro et al. used this model to study gene expressions following different stretch injury levels (Di Pietro et al., 2010). In their in vitro model of blast-induced TBI, Miller et al. exposed hippocampal brain slices to blast overpressures (Miller et al., 2015). For our study, we subjected the Organotypic Hippocampal Brain Slices (OHBS) to a focal mechanical trauma.
Following a well-established protocol for TBI-experiment (as described in Methods), we noticed that several factors might influence the results in such a set-up (Adamchik et al., 2000; Coburn et al., 2008; Grüßer et al., 2017; Krings et al., 2016; Loetscher et al., 2009; Roehl et al., 2010; Rossaint et al., 2009; Schoeler et al., 2012). Here, we describe a structured approach to generate more comparable results and discuss why specific eligibility criteria should be applied. We propose a six-step approach of comparing our results to ensure gaining more precise findings.
2. Methods
The data used for this study are derived from our TBI-experiments scrutinizing the impact of incubation with argon 50%, desflurane 6%, and their combination in 1459 OHBS. The final results of assessed interventions are presented and discussed in a separate paper (Grüßer et al., 2017). A specific approach to improve the quality of results in such a set-up was described in this study. This approach was approved by the local Institutional Ethics Review Committee and the Institute of Animal Research Blinded University Hospital’s animal protection representative, according to The German Animal Welfare Act (TierSchG § 4, III). We, Firstly, describe the well-established TBI-protocol briefly as described in the separate paper (Grüßer et al., 2017). Secondly, we outline the six steps developed to generate more comparable results (Grüßer et al., 2017).
2.1. Traumatic brain injury protocol
2.1.1. Organotypic hippocampal slices
The OHBS were obtained from 5-7-day-old mice pups (C57BL/6N, Charles River Laboratories, Sulzfeld, Germany) and kept in cultures as summarized in Suppl. 1, as previously described (Coburn et al., 2008; Grüßer et al., 2017; Krings et al., 2016; Loetscher et al., 2009; Roehl et al., 2010; Rossaint et al., 2009; Schoeler et al., 2012; Stoppini et al., 1991).
2.1.2. Traumatic brain injury
The experiments were performed as described before (Grüßer et al., 2017): the Growth Medium (GM) was exchanged by an Experimental Medium (EM) containing Propidium Iodide (PI). Then, baseline fluorescence images showing the cell damage before the experiment were taken and tissue plates were randomly assigned to their groups. Our experiment included four trauma and four non-trauma groups in which three different kinds of atmospheric interventions and one control (no intervention) were administered.
TBI was elicited in a similar way to the procedure described by other researchers (Adamchik et al., 2000; Coburn et al., 2008; Krings et al., 2016; Loetscher et al., 2009). A round stylus was positioned 7 mm above the hippocampus slice and then was dropped onto the slice positioned at a marked spot. After TBI was performed, EM was exchanged once more, and slices were incubated in the respective atmosphere. In non-TBI group, the slices underwent the same procedure except that no trauma was induced. After an incubation period of two hours, final fluorescence imaging was taken for all groups.
2.1.3. Microscopy and assessment of cell death
PI, a dye that rapidly enters cells with damaged membranes and binds to their DNA, was applied to assess the proportion of cell death (Macklis & Madison, 1990). Images were captured with a fluorescence microscope and MetaVue software (MetaVue, Molecular Devices, Sunnyvale, CA, USA). Using ImageJ software (National Institutes of Health, Bethesda, MD, USA), a histogram of the red channel was generated. The histogram listed the sum of all pixels sharing the same greyscale value from 0-255. Previous studies have shown that values below a threshold of 100 are caused by background fluorescence and only values above this threshold were summed up to assess the extent of the cell injury (Loetscher et al., 2009; Roehl et al., 2010; Schoeler et al., 2012). Hence, the sum of the pixels in an image represents the damage of a slice, as demonstrated in the separate paper (Grüßer et al., 2017).
2.1.4. Study statistics
SPSS V. 22, was used to calculate the Mean±SD error of the mean of the sum of pixels for each group. As a reference value, the trauma plus no intervention group’s mean value was normalized to unity. Statistical relevance was determined with the help of the Student t-test. P<0.05 were considered to be significant. The differences (∆) of the mean value of the sum of pixels for each intervention group before versus after a specific step are presented in percentage.
2.1.5. Recommendation: Six steps to generate more comparable results
Through a closer investigation of OHBS in our TBI-experiment, we identified several factors that might influence the results. To enhance comparability, we developed the following six steps, which are summarized in a model as shown in Figure 1, a flowchart which summarized the six-step approach as follow:
Brain cells have been studied in cultures for over 100 years (Ross Granville Harrison, 1910; Rose G. Harrison, Greenman, Mall, & Jackson, 1907; Hogue, 1952; Humpel, 2015; Millet & Gillette, 2012). Over time, different methods were developed to preserve thin brain slices and keep them viable for a longer time (Cho, Wood, & Bowlby, 2007; Gähwiler, 1981; Humpel, 2015; Stoppini, Buchs, & Muller, 1991). Slice cultures can be prepared from a variety of brain regions; however, hippocampal slice cultures are used most frequently and replicate many aspects of the in vivo state, making possible investigations into mechanisms of brain synapses (Bahr et al., 1995; Cho et al., 2007; Finley et al., 2004; Gähwiler, Capogna, Debanne, McKinney, & Thompson, 1997; Li, Han, & Wang, 2016; Noraberg et al., 2005).
Given that the hippocampal slice set-up also provides decent experimental access and allows for detailed regulation of the extracellular environment, it is not surprising that these systems have been widely used to study neurogenesis or to investigate diseases such as Alzheimer and stroke (Cho et al., 2007; Noraberg et al., 2005).
Several researchers utilized organotypic hippocampal slice cultures to scrutinize processes and possible treatment approaches for Traumatic Brain Injury (TBI) and described different methods in this regard (Adamchik, Frantseva, Weisspapir, Carlen, & Perez Velazquez, 2000; Coburn, Maze, & Franks, 2008; Di Pietro et al., 2010; Di Pietro et al., 2013; Harris et al., 2013; Hughes, Silva, Ahmed, Shreiber, & Morrison, 2014; Krings, Höllig, Grüsser, Rossaint, & Coburn, 2016; Loetscher et al., 2009; Miller et al., 2015; Morrison, Cater, Benham, & Sundstrom, 2006; Morrison et al., 2003; Roehl et al., 2010; Rossaint et al., 2009; Schoeler et al., 2012; Smith et al., 2016; Vogel et al., 2016).
Morrison et al. described an injury device that stretched the hippocampal brain slices equi-biaxially to simulate brain deformation through TBI (Morrison et al., 2006). Di Pietro et al. used this model to study gene expressions following different stretch injury levels (Di Pietro et al., 2010). In their in vitro model of blast-induced TBI, Miller et al. exposed hippocampal brain slices to blast overpressures (Miller et al., 2015). For our study, we subjected the Organotypic Hippocampal Brain Slices (OHBS) to a focal mechanical trauma.
Following a well-established protocol for TBI-experiment (as described in Methods), we noticed that several factors might influence the results in such a set-up (Adamchik et al., 2000; Coburn et al., 2008; Grüßer et al., 2017; Krings et al., 2016; Loetscher et al., 2009; Roehl et al., 2010; Rossaint et al., 2009; Schoeler et al., 2012). Here, we describe a structured approach to generate more comparable results and discuss why specific eligibility criteria should be applied. We propose a six-step approach of comparing our results to ensure gaining more precise findings.
2. Methods
The data used for this study are derived from our TBI-experiments scrutinizing the impact of incubation with argon 50%, desflurane 6%, and their combination in 1459 OHBS. The final results of assessed interventions are presented and discussed in a separate paper (Grüßer et al., 2017). A specific approach to improve the quality of results in such a set-up was described in this study. This approach was approved by the local Institutional Ethics Review Committee and the Institute of Animal Research Blinded University Hospital’s animal protection representative, according to The German Animal Welfare Act (TierSchG § 4, III). We, Firstly, describe the well-established TBI-protocol briefly as described in the separate paper (Grüßer et al., 2017). Secondly, we outline the six steps developed to generate more comparable results (Grüßer et al., 2017).
2.1. Traumatic brain injury protocol
2.1.1. Organotypic hippocampal slices
The OHBS were obtained from 5-7-day-old mice pups (C57BL/6N, Charles River Laboratories, Sulzfeld, Germany) and kept in cultures as summarized in Suppl. 1, as previously described (Coburn et al., 2008; Grüßer et al., 2017; Krings et al., 2016; Loetscher et al., 2009; Roehl et al., 2010; Rossaint et al., 2009; Schoeler et al., 2012; Stoppini et al., 1991).
2.1.2. Traumatic brain injury
The experiments were performed as described before (Grüßer et al., 2017): the Growth Medium (GM) was exchanged by an Experimental Medium (EM) containing Propidium Iodide (PI). Then, baseline fluorescence images showing the cell damage before the experiment were taken and tissue plates were randomly assigned to their groups. Our experiment included four trauma and four non-trauma groups in which three different kinds of atmospheric interventions and one control (no intervention) were administered.
TBI was elicited in a similar way to the procedure described by other researchers (Adamchik et al., 2000; Coburn et al., 2008; Krings et al., 2016; Loetscher et al., 2009). A round stylus was positioned 7 mm above the hippocampus slice and then was dropped onto the slice positioned at a marked spot. After TBI was performed, EM was exchanged once more, and slices were incubated in the respective atmosphere. In non-TBI group, the slices underwent the same procedure except that no trauma was induced. After an incubation period of two hours, final fluorescence imaging was taken for all groups.
2.1.3. Microscopy and assessment of cell death
PI, a dye that rapidly enters cells with damaged membranes and binds to their DNA, was applied to assess the proportion of cell death (Macklis & Madison, 1990). Images were captured with a fluorescence microscope and MetaVue software (MetaVue, Molecular Devices, Sunnyvale, CA, USA). Using ImageJ software (National Institutes of Health, Bethesda, MD, USA), a histogram of the red channel was generated. The histogram listed the sum of all pixels sharing the same greyscale value from 0-255. Previous studies have shown that values below a threshold of 100 are caused by background fluorescence and only values above this threshold were summed up to assess the extent of the cell injury (Loetscher et al., 2009; Roehl et al., 2010; Schoeler et al., 2012). Hence, the sum of the pixels in an image represents the damage of a slice, as demonstrated in the separate paper (Grüßer et al., 2017).
2.1.4. Study statistics
SPSS V. 22, was used to calculate the Mean±SD error of the mean of the sum of pixels for each group. As a reference value, the trauma plus no intervention group’s mean value was normalized to unity. Statistical relevance was determined with the help of the Student t-test. P<0.05 were considered to be significant. The differences (∆) of the mean value of the sum of pixels for each intervention group before versus after a specific step are presented in percentage.
2.1.5. Recommendation: Six steps to generate more comparable results
Through a closer investigation of OHBS in our TBI-experiment, we identified several factors that might influence the results. To enhance comparability, we developed the following six steps, which are summarized in a model as shown in Figure 1, a flowchart which summarized the six-step approach as follow:
2.1.5.1. Confirmation
In the first step, it is recommended to verify that each picture taken shows only one slice, amounts to the pixels of this slice, and thus fully shows its cell damage. We suggest enhancing the contrast of each image captured (only for better visibility, pixel counts are determined as explained above) (Figure 2.1). This procedure allows for the identification of a second slice in the picture and eventually the blackening of this area. Further, a thorough evaluation of each photograph is advantageous to detect small PI-clots or fusels. Finally, we are sure that each image taken before the experiment is matched with the correct one at later time-points of assessment (Figure 2.1, Example image step 1).
2.1.5.2. Identification of pre-damaged slices at baseline assessment and exclusion
Other researchers have pointed to the fact, that they excluded slices from their experiments if they were damaged at baseline assessment or showed high levels of PI fluorescence (Adamchik et al., 2000; Harris et al., 2013; Krings et al., 2016; Miller et al., 2015). However, no details were provided, and in many papers, we did not find any information that slices were excluded. As inserts are moved for medium changes and are exposed to temperature fluctuations, pre-damage to the slice at baseline assessment is a common event. Moreover, preparation techniques and timing may still improve within the course of the experiment.
It is crucial to avoid pre-damaged slices entering into the experiment, as their trauma intensity may falsify the results after the incubation time. Since it is unknown whether a higher amount of injured cells at baseline assessment affect or even exponentially increase the development of secondary injury, only minimally pre-damaged slices should be considered in the experiment. An objective threshold should be defined to avoid bias in identification and outsourcing of these slices. In our research, if baseline image histograms presented more than 1000 pixels, they would be sorted out as they were considered to be pre-damaged (Figure 2.2, Example image of step 2).
2.1.5.3. Preparation margin error
Motor skills develop over training sessions and probably through different stages (Luft & Buitrago, 2005). Similarly, preparation techniques improve when regularly dissecting the hippocampi. A typical feature we observed, especially within the first rounds of prepared slices, was a prominent margin due to-as suggested here-errors in preparation. Even though preparation steps were always strictly followed, some of the slices presented with noticeable frames of PI-detectable cells. The pictures taken at baseline assessment did not necessarily exceed an integral of >1000 pixels.
After an incubation period of two hours, however, PI fluorescence was highly visible, and thus slices with such a prominent margin distorted the outcome of the affected groups. In order to objectively decide on an allowable error in preparation and an average damage which would inevitably occur, the following threshold was applied: if the PI-detectable cell death in a margin exceeded >250 pixels, this margin was considered to be prominent enough to be an error stemming from errors in the preparation and its pixels were not added to the full integration of pixels in this picture (Figure 2.3). We took the same approach for slices presenting with a “one-celled” margin, which was observed in only two experimental series (Figure 2.3, Example image of step 3).
2.1.5.4. Morphological peculiarities
In this study, we noticed several morphological peculiarities in varying degrees. These peculiarities include what we describe as “holes”, “frayed margins”, and “emigrated cells” (Figure 2.4). A closer investigation of these phenomena is suggested for future studies, as we do not know why they occurred and cannot evaluate whether electrophysiological consequences are entailed. We observed that “holes” often went in line with “frayed margins” and “emigrated cells”.
However, these features also occurred independently. Additionally, by enhancing the contrast of the image, we noticed that emigrated cells even existed at baseline assessment, but this was not always detectable by PI. Excluding all slices with a specific morphology would significantly decrease the number of slices available in particular groups. Therefore, we decided to refer to step 1: if slices with peculiarities were presented with <1000 pixels at baseline assessment, they would be taken into consideration for the experiment (Figure 2.4, Example image of step 4).
2.1.5.5. Variation of trauma intensities
During the experiment, we observed that trauma intensities varied over time and that some experimental series, even within one group presented with different trauma severities despite the same dropping height was ensured (Figure 2.5). Thus, we saw the necessity to define a scope of trauma strength. A boxplot analysis was performed for the trauma control group, and extreme outliers were identified (n=2) and excluded. The new mean value of the trauma control group plus two times the standard deviation served as an upper threshold for all the other trauma groups.
Concerning trauma intensities and depending on where the pin hits the slice, more or less tissue can be injured by secondary injury; that is, if the pin hits the slice close to the margin there is less hippocampus to be affected in the following time. Also the shapes of the hippocampi- as we experienced- vary where we found it is unlikely to work with remarkably similar contoured slices. Hence, our approach was to consider all slices except for those that were not hit by the pin (n=7).
In our experiment, we identified several borderline cases, mostly in the intervention group presenting with the least secondary injury, where it appeared to be difficult to make a clear decision. Future researchers are recommended to set a defined size before the experiment that has to be considered for inclusion criteria, including the hippocampus itself (which is also of importance for the non-trauma groups), the pin’s impression, and the area of secondary injury (Figure 2.5, Example image for step 5).
2.1.5.6. Susceptibility of non-trauma groups
We noticed that it is crucial to adjust the focus of the camera in taking pictures, particularly in the non-trauma groups, where trauma intensities are minimal. Blurry images can present with a lower amount of greyscale pixels as the majority of pixels are evaluated as background fluorescence. Hence, in a few cases, pre-damaged slices may not be excluded by the 1000-pixels threshold and then present with excessive greyscale values after the incubation time. Also- the other way round- a misleadingly low number of pixels is possible when the picture was taken after the incubation time is blurry.
To prevent biases by manually evaluating the sharpness of hundreds of pictures, we first excluded all images in the non-trauma groups that had lower pixel numbers after the incubation period than at baseline assessment. We subjectively excluded 11 slices due to their size as, for example, they were too small to be considered a hippocampal slice. In the future, a defined size of the hippocampus should be set before the experiment. In the next step, we performed a boxplot-analysis in each non-trauma group to exclude outliers. Thus, we objectively excluded two kinds of slices: the slices presenting with excessive trauma (as the pictures showing the pre-damage at baseline assessment were blurry), and the slices damaged due to careless handling or temperature change (Figure 2-6, Example image of step 6).
3. Results
We proposed a six-step approach to enhance comparability and applied the proposed model to the data derived from our experiments on 1459 OHBS, as mentioned above (Figure 1). Then we presented exemplary results that demonstrated how the outcome could differ when a measure was applied vs. when it was not.
After the first checkpoint (step 1) (confirmation), 1459 slices were assessed for pre-damage (step 2). As Figure 3.1 shows, the difference between two non-trauma groups seems to be significant at first glance; yet, when it is ascertained that baseline slices in both groups were presented with similar PI-detectable trauma intensities, this difference did not turn out to be significant anymore (P=0.004 vs. P=0.716; ∆ intervention 2: 0.27; ∆ intervention 3: 0.56). In the next step, the slices were screened for preparatory margins (step 3). Figure 3.2 demonstrates the difference in results when preparation errors are considered vs. when they are not. Misleadingly, the non-trauma/intervention group 2 would have been considered significantly more injured than the non-trauma/intervention group 1 (P=0.004 vs. P=0.444; ∆ intervention 1: 0.02 ∆ intervention 2: 0.24).
Without applying this measure, a difference between groups would have mistakenly been attributed to the specific interventions. By subtracting margins stemming from errors in the preparatory process, more slices met the baseline assessment threshold and were considered for the next steps (393 slices after step 2 vs. 399 slices after step 3 in the trauma groups and 568 vs. 637 slices in the non-trauma groups). Morphological peculiarities, as described in step 4 require further investigation. Step 5 aims to set a scope concerning trauma intensities in the trauma groups.
According to Figure 3.3, without a set scope, the trauma/intervention group 3 would misleadingly have been considered to generate a neuro-harming effect in comparison to the trauma/intervention group 2 (P<0.001 vs. P=0.304; ∆ intervention 2: 0.02; ∆ intervention 3: 0.31). Identifying and excluding outliers in the non-trauma groups (step 6) revealed that the difference between the non-trauma/no intervention group and the non-trauma/intervention group 3 was not significant (P=0.016 vs. P=0.075; ∆ no intervention: 0.24; ∆ intervention 3: 0.40) (Figure 3.4).
After the six-step approach, the number of slices was narrowed down to 547 pictures in the non-trauma groups and 341 photos in the trauma groups (Figure 1). Therefore, 61% of the slices met the proposed inclusion criteria (Figure 3; results demonstrating more precise findings).
4. Discussion
Working with OHBS in a TBI-experiment, we noticed that several factors might influence the results. To our knowledge, detailed objective manners to generate more comparable results in such a susceptible set-up have not been discussed so far. The proposed model aims to set precise requirements to lower the possibility of pre-damaged slices entering into the experiment and outliers twisting the final results. This methodological approach provides the first framework to cope with the broad variation observed in these models. Our exemplary results show that more precise findings are ensured by setting equal requirements for all slices.
In vitro models provide a useful tool to scrutinize processes and treatment options in the initial step. Between dissociated cell cultures and the in vivo state, OHBS presents a potent alternative (Humpel, 2015; Li et al., 2016; Morrison, Saatman, Meaney, & McIntosh, 1998; Noraberg et al., 2005). The hippocampal circuits in organotypic brain slices, which are usually derived from early postnatal animals, are found to be similar to the in vivo state; however, some differences have been detected (Gähwiler et al., 1997; Gogolla, Galimberti, DePaola, & Caroni, 2006).
Of course, variables like perfusion, intracranial pressure, or the blood-brain barrier have not yet reflected in organotypic brain slice models, and the use of adult donor animals for organotypic brain slices has been scarce (Humpel, 2015; Li et al., 2016). Nonetheless, slices have remained 100-150 µm thick with a three-dimensional organization of connectivity (Gähwiler et al., 1997; Gogolla et al., 2006; Li et al., 2016; Stoppini et al., 1991). Notably, outcomes of studies working with OHBS have successfully been reproduced in in vivo models demonstrating the potential of this setup. For example, argon and xenon displayed a neuroprotective profile in OHBS models which could be substantiated in in vivo studies (Brücken et al., 2013; Coburn et al., 2008; Harris et al., 2013; Loetscher et al., 2009; Zhuang et al., 2012).
An important implication of research with organotypic brain slices is the reduction of the number of animal experiments (Humpel, 2015; Li et al., 2016). Further, in in vitro models of CNS injury, the associated costs are lower (Morrison et al., 1998). In their reviews on brain slice models, Noraberg et al. concluded that organotypic brain slices emerged to be an increasingly popular tool to investigate neurological diseases. Cho et al. asserted that through advancements of brain slices and technological innovations in this field now a tremendous potential exists to address questions quicker (Cho et al., 2007; Noraberg et al., 2005).
To our knowledge, precise and objective procedures to make results more comparable within one experiment and among different studies have not been discussed in detail yet. Retrospectively, Guy et al. method of determining slice thickness as a parameter for OHBS health might present a useful additional measure (Guy, Rupert, Sandberg, & Weber, 2011). In their review, Humpel et al. generally recommended withdrawing thick and not flattened slices. An outgrowth of cells from the edge of the living slices and a change in color to transparent grey color in the first week are criteria described for ‘good’ slices (Humpel, 2015).
De Simoni et al. pointed the difficulty to identify pyramidal cell layers in an unhealthy slice (De Simoni & Yu, 2006). Also, Gogolla et al. recommended not to culture slices with a lesioned CA1 region or a lesioned/detached dentate gyrus but only slices ‘with nice cell layers in the dentate gyrus and CA1-3 and smooth margins’ seems to be a reasonable approach to be considered for future researchers (Gogolla et al., 2006). Our model mainly relied on PI-detectable cell death, but other cell-specific markers than PI can be used (Humpel, 2015).
Experimental papers often do not report about slice selection. Some researchers have pointed to difficulties as variation in PI-uptake, incomplete layers or pre-damage, but did not explain objective criteria concerning how or why brain slices were excluded or included (Adamchik et al., 2000; Harris et al., 2013; Krings et al., 2016; Li et al., 2016; Miller et al., 2015). For clinical trials, participants must meet specific standards, fulfill inclusion criteria, or be prohibited from participating due to exclusion criteria (National Institutes of Health, 2016). This method neither protects the participants’ safety nor ensures that the subsequent results are more comparable as the study population is more alike.
Within our TBI-experiment in OHBS, we found that results differ when hippocampal brain slices undergo specific assessments for eligibility and that hippocampal brain slices can present with pre-damage or specific peculiarities and identified factors that might affect the outcome.
In general, researchers should work as cautiously as possible to avoid imprecise preparation, temperature changes for a long time, accidental shaking, or changes in the experimental process. Although all precautions may be observed, organizational challenges concerning timing, the microscope setting, or the medium may occur. Fluctuations in the severity of trauma may distort the outcome. Although the reasons are not clear, our results show that hippocampal brain slices do not necessarily present an unconditionally homogenous group. Thus, eligibility criteria for OHBS present a plausible conclusion.
To avoid bias by manually evaluating slices, the criteria set should be as objective as possible. If slices are uniform and interventions are consistent, the eligibility criteria do not interfere with the outcome. If any inconsistencies occur, the applied measures increase the chances of more precise conclusions to be drawn. In our experiment, for example, a considerable amount of data were estimated unusable as the slices were considered pre-damaged. Since this threshold was chosen arbitrarily, we could thus ensure that equal requirements were set for all experimental series.
Our results demonstrate that outcomes in a TBI experiment with OHBS should be critically analyzed as inhomogeneity may occur. Altogether, the proposed six steps are designed to consider possible errors; they also help to contextualize the findings and make results more comparable. In the future, enhanced comparability may decrease the number of animals needed.
This framework presents a first attempt to enhance comparability in an in vitro model of TBI in OHBS. Further research is recommended to confirm and further develop this model in similar studies. It might improve the quality of the results if the general recommendations listed above were specified and also considered in the proposed approach because of PI-detectable cell death, and the proposed measures might not be sufficient.
Some challenges that could not be adequately addressed would remain. First, the proposed thresholds are chosen arbitrarily, and future researchers are recommended to find an analytical solution of what should be considered ineligible, i.e. pre-damage.
Another disadvantage might be the visible preparatory margins with less than 250 pixels could still enter the experiment. As the margins had to be identified and blackened manually, this represents a subjective component and remains a potential source of error. We aimed to blacken only areas of cell death, possibly stemming from errors in the preparation process as the morphological peculiarities and their origin are unknown. However, it is not always clear whether a specific area should be considered as a “preparatory error”, a “frayed margin”, a “hole”, or as “emigrated cells”. For further studies, we would also recommend to regularly allocate prepared slices in one series to all groups to distribute potential errors equally.
Despite all measures taken, the slices that were too small to be considered as a hippocampus or some slices which were not hit by the pin had to be identified. We tried to minimize the number of slices excluded manually, so this problem presented a subjective choice, and we recommend defining inclusion criteria concerning the size of the hippocampi and the trauma area for future research. This suggestion seems particularly useful, as the trauma produced by the pin in our experiment, does not always resemble a proper circle, as are observable in images shown in other papers (Adamchik et al., 2000; Roehl et al., 2010). It is unclear whether this is typical as the injury spreads non-circularly throughout the slice depending on differences in susceptibility of cells or is caused by other factors, as an oblique fall.
Although the framework was constructed to avoid future errors, there may be cases that were not fixed. For example, our measures to exclude the most blurry pictures might not be sufficient to identify them. We included all slices and applied the measures proposed; however, numerical thresholds, i.e. morphological completeness, cannot be identified. Nevertheless, we sorted out a considerable amount of data and when the inclusion criteria failed to meet. This model presented a first approach to make the research with hippocampal brain slices in an in vitro model of TBI more comparable.
5. Conclusion
Several factors may influence experimental outcomes in a TBI-experiment in OHBS. To achieve the most precise result, we should set checkpoints, define pre-damaged slices, and restrain from entering into the experiment. Besides, we should detect unrealistic outliers and eventually exclude them. We presented objective approaches in our framework and included thresholds and boxplot-analyses. Our exemplary results demonstrate more comparable findings. We recommend future researchers to confirm and develop further this model and to scrutinize morphological peculiarities, their origin, and their potential impact on cell survival and electrophysiological functioning.
Ethical Considerations
Compliance with ethical guidelines
All ethical principles were considered in this article.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Authors' contributions
All authors contributed in preparing this article.
Conflict of interest
The authors declared no conflict of interest.
Acknowledgments
We thank Prof. Joachim Weis, head of the Department of Neuropathology, RWTH Aachen University Hospital, Aachen, Germany, and his team for their support in the culturing of the slices.
References
- Adamchik, Y., Frantseva, M. V., Weisspapir, M., Carlen, P. L., & Velazquez, J. P. (2000). Methods to induce primary and secondary traumatic damage in organotypic hippocampal slice cultures. Brain Research Protocols, 5(2), 153-8. [DOI:10.1016/S1385-299X(00)00007-6]
- Bahr, B. A., Kessler, M., Rivera, S., Vanderklish, P. W., Hall, R. A., Mutneja, M. S., et al. (1995). Stable maintenance of glutamate receptors and other synaptic components in long-term hippocampal slices. Hippocampus, 5(5), 425-39. [DOI:10.1002/hipo.450050505] [PMID]
- Brücken, A., Cizen, A., Fera, C., Meinhardt, A., Weis, J., Nolte, K., et al. (2013). Argon reduces neurohistopathological damage and preserves functional recovery after cardiac arrest in rats. British Journal of Anaesthesia, 110(suppl. 1), i106-i12. [DOI:10.1093/bja/aes509] [PMID]
- Cho, S., Wood, A., & Bowlby, M. R. (2007). Brain slices as models for neurodegenerative disease and screening platforms to identify novel therapeutics. Current Neuropharmacology, 5(1), 19-33. [DOI:10.2174/157015907780077105] [PMID] [PMCID]
- Coburn, M., Maze, M., & Franks, N. P. (2008). The neuroprotective effects of xenon and helium in an in vitro model of traumatic brain injury. Critical Care Medicine, 36(2), 588-95. [DOI:10.1097/01.CCM.0B013E3181611F8A6] [PMID]
- De Simoni, A., & Lily, M. Y. (2006). Preparation of organotypic hippocampal slice cultures: Interface method. Nature Protocols, 1(3), 1439-45. [DOI:10.1038/nprot.2006.228] [PMID]
- Di Pietro, V., Amin, D., Pernagallo, S., Lazzarino, G., Tavazzi, B., Vagnozzi, R., et al. (2010). Transcriptomics of traumatic brain injury: gene expression and molecular pathways of different grades of insult in a rat organotypic hippocampal culture model. Journal of Neurotrauma, 27(2), 349-59. [DOI:10.1089/neu.2009.1095] [PMID]
- Di Pietro, V., Amorini, A. M., Tavazzi, B., Hovda, D. A., Signoretti, S., Giza, C. C., et al. (2013). Potentially neuroprotective gene modulation in an in vitro model of mild traumatic brain injury. Molecular and Cellular Biochemistry, 375(1-2), 185-98. [DOI:10.1007/s11010-012-1541-2] [PMID]
- Finley, M., Fairman, D., Liu, D., Li, P., Wood, A., & Cho, S. (2004). Functional validation of adult hippocampal organotypic cultures as an in vitro model of brain injury. Brain Research, 1001(1-2), 125-32. [DOI:10.1016/j.brainres.2003.12.009] [PMID]
- Gähwiler, B. H. (1981). Organotypic monolayer cultures of nervous tissue. Journal of Neuroscience Methods, 4(4), 329-42. [DOI:10.1016/0165-0270(81)90003-0]
- Gähwiler, B. H., Capogna, M., Debanne, D., McKinney, R. A., & Thompson, S. M. (1997). Organotypic slice cultures: A technique has come of age. Trends in Neurosciences, 20(10), 471-7. [DOI:10.1016/S0166-2236(97)01122-3]
- Gogolla, N., Galimberti, I., DePaola, V., & Caroni, P. (2006). Preparation of organotypic hippocampal slice cultures for long-term live imaging. Nature protocols, 1(3), 1165-71. [DOI:10.1038/nprot.2006.168] [PMID]
- Grüßer, L., Blaumeiser-Debarry, R., Krings, M., Kremer, B., Höllig, A., Rossaint, R., et al. (2017). Argon attenuates the emergence of secondary injury after traumatic brain injury within a 2-hour incubation period compared to desflurane: An in vitro study. Medical Gas Research, 7(2), 93-100. [DOI:10.4103/2045-9912.208512] [PMID] [PMCID]
- Guy, Y., Rupert, A. E., Sandberg, M., & Weber, S. G. (2011). A simple method for measuring organotypic tissue slice culture thickness. Journal of Neuroscience Methods, 199(1), 78-81. [DOI:10.1016/j.jneumeth.2011.03.027] [PMID] [PMCID]
- Harris, K., Armstrong, S. P., Campos-Pires, R., Kiru, L., Franks, N. P., & Dickinson, R. (2013). Neuroprotection against traumatic brain injury by xenon, but not argon, is mediated by inhibition at the N-methyl-D-aspartate receptor glycine site. Anesthesiology, 119(5), 1137-48. [DOI:10.1097/ALN.0b013e3182a2a265] [PMID]
- Harrison, R. G. (1910). The outgrowth of the nerve fiber as a mode of protoplasmic movement. Journal of Experimental Zoology, 9(4), 787-846. [DOI:10.1002/jez.1400090405]
- Harrison, R. G., Greenman, M. J., Mall, F. P., & Jackson, C. M. (1907). Observations of the living developing nerve fiber. The Anatomical Record, 1(5), 116-28. [DOI:10.1002/ar.1090010503]
- Hogue, M. J. (1952). Review of studies of human fetal brain cells in tissue cultures. Etudes Neonatales, 1(2), 1-13. [DOI:10.1016/0014-4827(52)90129-8]
- Hughes, R. H., Silva, V. A., Ahmed, I., Shreiber, D. I., & Morrison III, B. (2014). Neuroprotection by genipin against reactive oxygen and reactive nitrogen species-mediated injury in organotypic hippocampal slice cultures. Brain Research, 1543, 308-14. [DOI:10.1016/j.brainres.2013.11.020] [PMID]
- Humpel, C. (2015). Organotypic brain slice cultures: A review. Neuroscience, 305, 86-98. [DOI:10.1016/j.neuroscience.2015.07.086] [PMID] [PMCID]
- Krings, M., Höllig, A., Grüsser, L., Rossaint, R., & Coburn, M. (2016). Desflurane impairs outcome of organotypic hippocampal slices in an in vitro model of traumatic brain injury. Medical Gas Research, 6(1), 3-9. [DOI:10.4103/2045-9912.179338] [PMID] [PMCID]
- Li, Q., Han, X., & Wang, J. (2016). Organotypic hippocampal slices as models for stroke and traumatic brain injury. Molecular Neurobiology, 53(6), 4226-37. [DOI:10.1007/s12035-015-9362-4] [PMID] [PMCID]
- Loetscher, P. D., Rossaint, J., Rossaint, R., Weis, J., Fries, M., Fahlenkamp, A., et al. (2009). Argon: Neuroprotection in in vitro models of cerebral ischemia and traumatic brain injury. Critical Care, 13(6), R206. [DOI:10.1186/cc8214] [PMID] [PMCID]
- Luft, A. R., & Buitrago, M. M. (2005). Stages of motor skill learning. Molecular Neurobiology, 32(3), 205-16. [DOI:10.1385/MN:32:3:205]
- Macklis, J. D., & Madison, R. D. (1990). Progressive incorporation of propidium iodide in cultured mouse neurons correlates with declining electrophysiological status: A fluorescence scale of membrane integrity. Journal of Neuroscience Methods, 31(1), 43-46. [DOI:10.1016/0165-0270(90)90007-3]
- Miller, A. P., Shah, A. S., Aperi, B. V., Budde, M. D., Pintar, F. A., Tarima, S., et al. (2015). Effects of blast overpressure on neurons and glial cells in rat organotypic hippocampal slice cultures. Frontiers in Neurology, 6(20), 1-20. [DOI:10.3389/fneur.2015.00020]
- Millet, L. J., & Gillette, M. U. (2012). Over a century of neuron culture: From the hanging drop to microfluidic devices. The Yale Journal of Biology and Medicine, 85(4), 501-21. [PMID] [PMCID]
- Morrison Iii, B. A., Saatman, K. E., Meaney, D. F., & Mcintosh, T. K. (1998). In vitro central nervous system models of mechanically induced trauma: A review. Journal of Neurotrauma, 15(11), 911-28. [DOI:10.1089/neu.1998.15.911] [PMID]
- Morrison III, B., Cater, H. L., Benham, C. D., & Sundstrom, L. E. (2006). An in vitro model of traumatic brain injury utilising two-dimensional stretch of organotypic hippocampal slice cultures. Journal of Neuroscience Methods, 150(2), 192-201. [DOI:10.1016/j.jneumeth.2005.06.014] [PMID]
- Morrison, B., Cater, H. L., Wang, C. C., Thomas, F. C., Hung, C. T., Ateshian, G. A., & Sundstrom, L. E. (2003). A tissue level tolerance criterion for living brain developed with an in vitro model of traumatic mechanical loading. Stapp Car Crash Journal, 47, 93-105. [DOI:10.4271/2003-22-0006] [PMID]
- National Institutes of Health. (2016). Learn about clinical studies. Retrieved from https://clinicaltrials.gov/ct2/about-studies/learn#WhoCanParticipate
- Noraberg, J., Poulsen, F. R., Blaabjerg, M., Kristensen, B. W., Bonde, C., Montero, M., et al. (2005). Organotypic hippocampal slice cultures for studies of brain damage, neuroprotection and neurorepair. Current Drug Targets-CNS & Neurological Disorders, 4(4), 435-52. [DOI:10.2174/1568007054546108] [PMID]
- Roehl, A. B., Hein, M., Loetscher, P. D., Rossaint, J., Weis, J., Rossaint, R., et al. (2010). Neuroprotective properties of levosimendan in an in vitro model of traumatic brain injury. BMC Neurology, 10, 97. [DOI:10.1186/1471-2377-10-97] [PMID] [PMCID]
- Rossaint, J., Rossaint, R., Weis, J., Fries, M., Rex, S., & Coburn, M. (2009). Propofol: Neuroprotection in an in vitro model of traumatic brain injury. Critical Care, 13, R61. [DOI:10.1186/cc7795] [PMID] [PMCID]
- Schoeler, M., Loetscher, P. D., Rossaint, R., Fahlenkamp, A. V., Eberhardt, G., Rex, S., et al. (2012). Dexmedetomidine is neuroprotective in an in vitro model for traumatic brain injury. BMC Neurology, 12, 20. [DOI:10.1186/1471-2377-12-20] [PMID] [PMCID]
- Smith, M., Piehler, T., Benjamin, R., Farizatto, K. L., Pait, M. C., Almeida, M. F., et al. (2016). Blast waves from detonated military explosive reduce GluR1 and synaptophysin levels in hippocampal slice cultures. Experimental Neurology, 286, 107-15. [DOI:10.1016/j.expneurol.2016.10.002] [PMID] [PMCID]
- Stoppini, L., Buchs, P. A., & Muller, D. (1991). A simple method for organotypic cultures of nervous tissue. Journal of Neuroscience Methods, 37(2), 173-82. [DOI:10.1016/0165-0270(91)90128-M]
- Vogel III, E. W., Effgen, G. B., Patel, T. P., Meaney, D. F., Bass, C. R. D., & Morrison III, B. (2016). Isolated primary blast inhibits long-term potentiation in organotypic hippocampal slice cultures. Journal of Neurotrauma, 33(7), 652-61. [DOI:10.1089/neu.2015.4045] [PMID] [PMCID]
- Zhuang, L., Yang, T., Zhao, H., Fidalgo, A. R., Vizcaychipi, M. P., Sanders, R. D., et al. (2012). The protective profile of argon, helium, and xenon in a model of neonatal asphyxia in rats. Critical Care Medicine, 40(6), 1724-30. [DOI:10.1097/CCM.0b013e3182452164] [PMID]
Type of Study: Original |
Subject:
Cellular and molecular Neuroscience
Received: 2017/09/7 | Accepted: 2018/09/22 | Published: 2019/09/1
Received: 2017/09/7 | Accepted: 2018/09/22 | Published: 2019/09/1
Send email to the article author
| Rights and permissions | |
 |
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License. |





.PNG)

